Institute of Bioinformatics Münster
Sorted Blast Graph - Gui
Usage
After downloading the application and start with
java -jar /path/to/sbg.jarin a terminal.
Input
This application takes a set of blast output of the following format
| Column | Name |
|---|---|
| 0 | query name |
| 1 | subject name |
| 2 | percent identities |
| 3 | aligned length |
| 4 | number of mismatched positions |
| 5 | number of gap positions |
| 6 | query sequence start |
| 7 | query sequence end |
| 8 | subject sequence start |
| 9 | subject sequence end |
| 10 | e-value |
| 11 | bit score |
Example
SAM50_AMEBA SAM50_AMEBA 100.00 118 0 0 1 118 1 118 9e-67 243 SAM50_AMEBA gi|126338997|ref|XP_001362470.1| 29.66 118 70 4 3 115 151 260 4e-09 52.4 SAM50_AMEBA gi|114686851|ref|XP_001172164.1| 28.81 118 71 4 3 115 72 181 1e-08 50.8
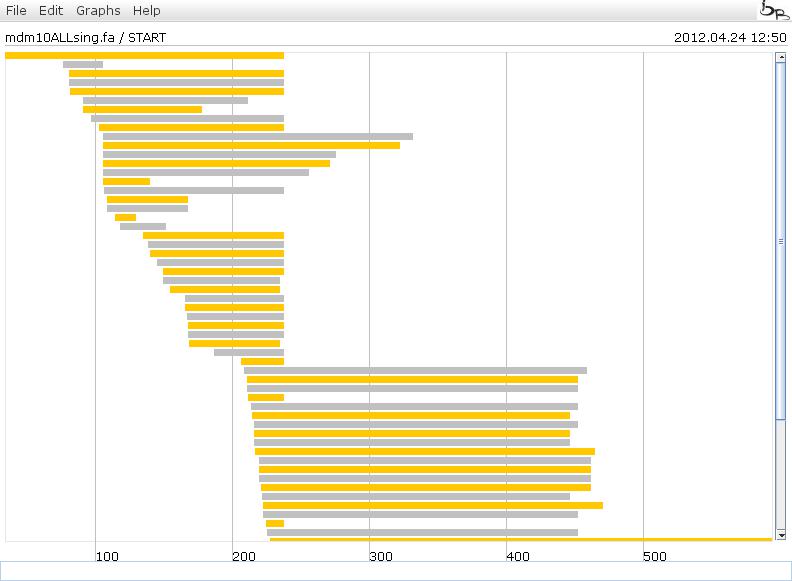
Generated Graphs
The generated lines are ordered in 3 different ways. The lines have the information about the sequence, the start and end positions where the fragment was found.
2025-06-05 09:23